MetaboT
Multi-agent LLM framework for querying mass spectrometry metabolomics knowledge graphs in natural language
MetaboT¶
MetaboT helps researchers ask natural-language questions over metabolomics knowledge graphs and receive answers backed by executable SPARQL queries. The system combines schema-aware prompting, multi-agent orchestration, entity resolution against authoritative resources, and optional interpretation of results.
The public demonstrator is available at metabot.holobiomicslab.eu, and the default local setup targets the ENPKG endpoint built from an open dataset of 1,600 plant extracts.
Why MetaboT?¶
- It translates natural-language metabolomics questions into executable SPARQL.
- It reduces hallucinations by resolving taxa, targets, chemical classes, and structures before query generation.
- It exposes a transparent, inspectable workflow instead of a single opaque prompt.
- It can be run from the command line, through Streamlit, or in Docker.
Validation Snapshot¶
The latest manuscript reports the following ENPKG benchmark results:
| System | Overall accuracy | High-complexity accuracy |
|---|---|---|
| GPT-4o single-shot | 8.16% | 0.00% |
| MetaboT with GPT-4o mini | 12.24% | 15.79% |
| MetaboT with GPT-4o | 83.67% | 78.95% |
These scores are reported over 49 scored questions from a 50-question benchmark, after excluding one refinement artifact discussed in the manuscript.
Architecture Overview¶
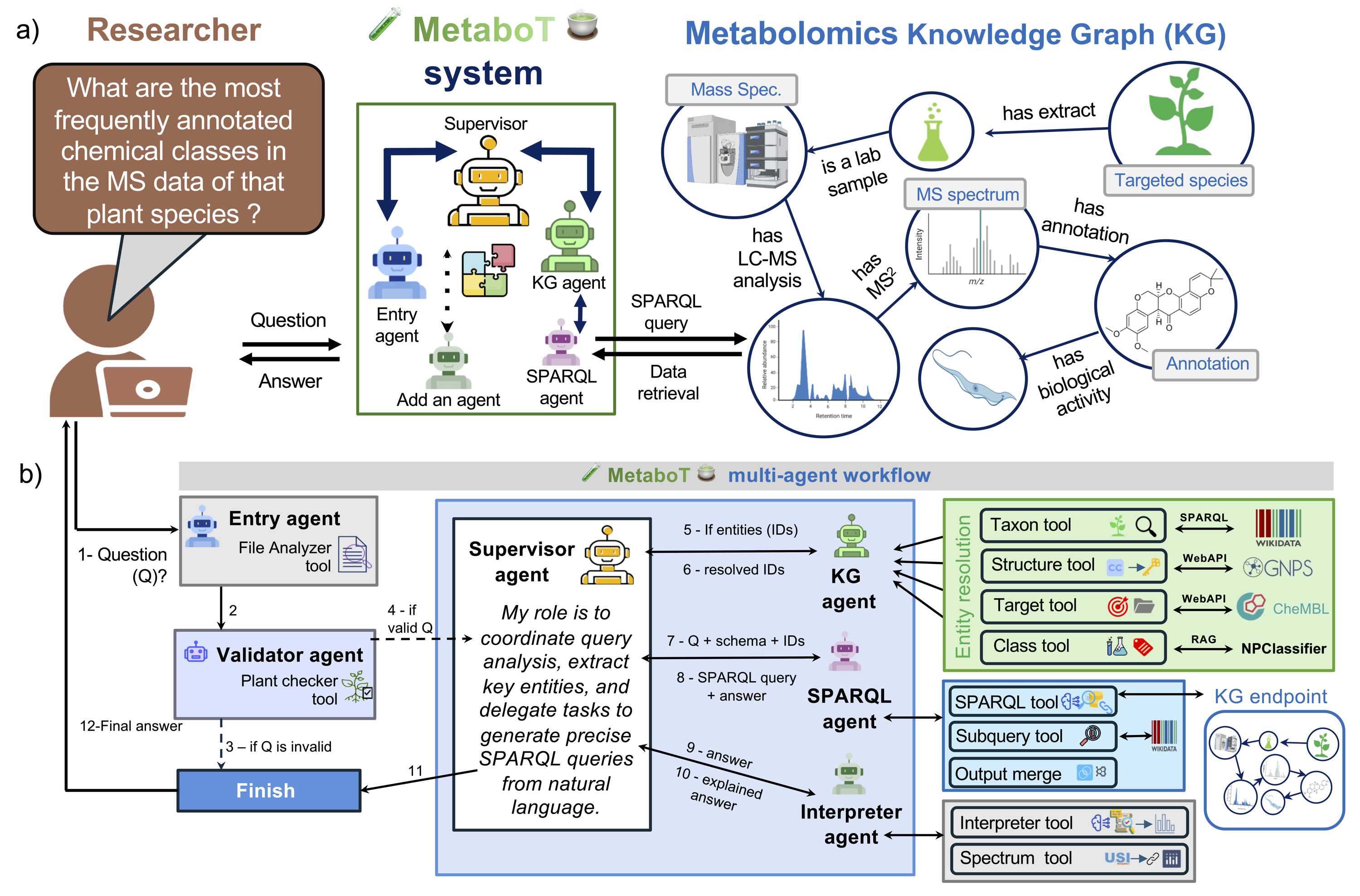
MetaboT orchestrates six main roles:
Entry Agentdecides whether the user is asking a new knowledge question or a follow-up.Validator Agentchecks whether the question matches the graph's schema and available data.Supervisor Agentroutes the request through the workflow.KG Agentresolves entities using tools connected to resources such as Wikidata, ChEMBL, NPClassifier, and GNPS.SPARQL Query Runner Agentbuilds and executes the query throughGraphSparqlQAChain.Interpreter Agentsummarizes the result and can generate plots when requested.
In the current codebase, the manuscript's KG Agent role is implemented by ENPKG_agent.
Workflow at a Glance¶
```mermaid graph TD A[User question] → B[Entry Agent] B → C[Validator Agent] C → D[Supervisor Agent] D → E[ENPKG_agent / KG Agent] D → F[SPARQL Query Runner Agent] F → G[Knowledge graph endpoint] D → H[Interpreter Agent] E → D F → D H → D D → I[Answer, SPARQL and CSV]
```
Quick Links¶
Citation¶
If you use MetaboT, please cite the current manuscript:
MetaboT: An LLM-based Multi-Agent Framework for Interactive Analysis of Mass Spectrometry Metabolomics Knowledge Graphs Research Square preprint. DOI: 10.21203/rs.3.rs-6591884/v1
The benchmark release and archived evaluated version are available on Zenodo.